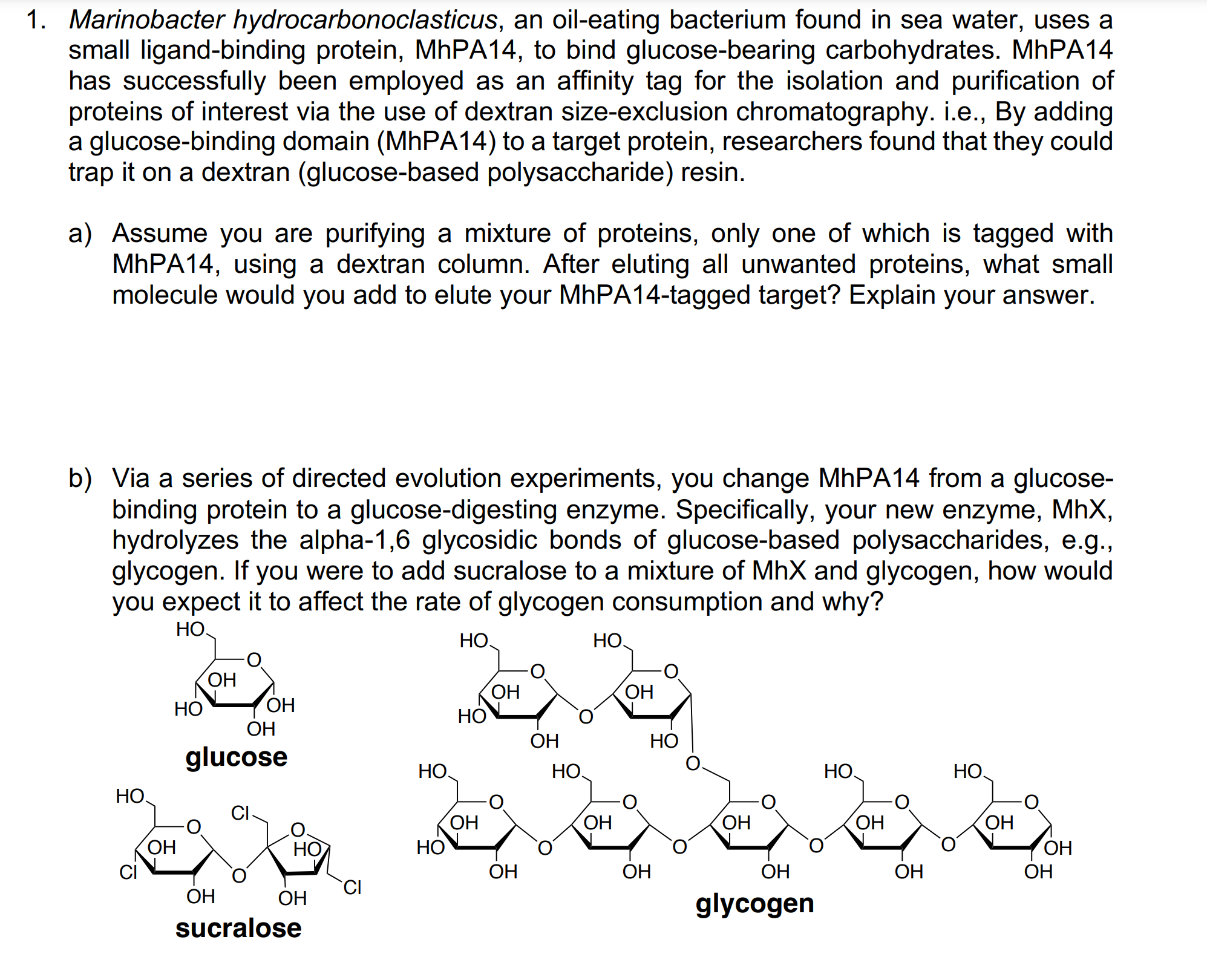
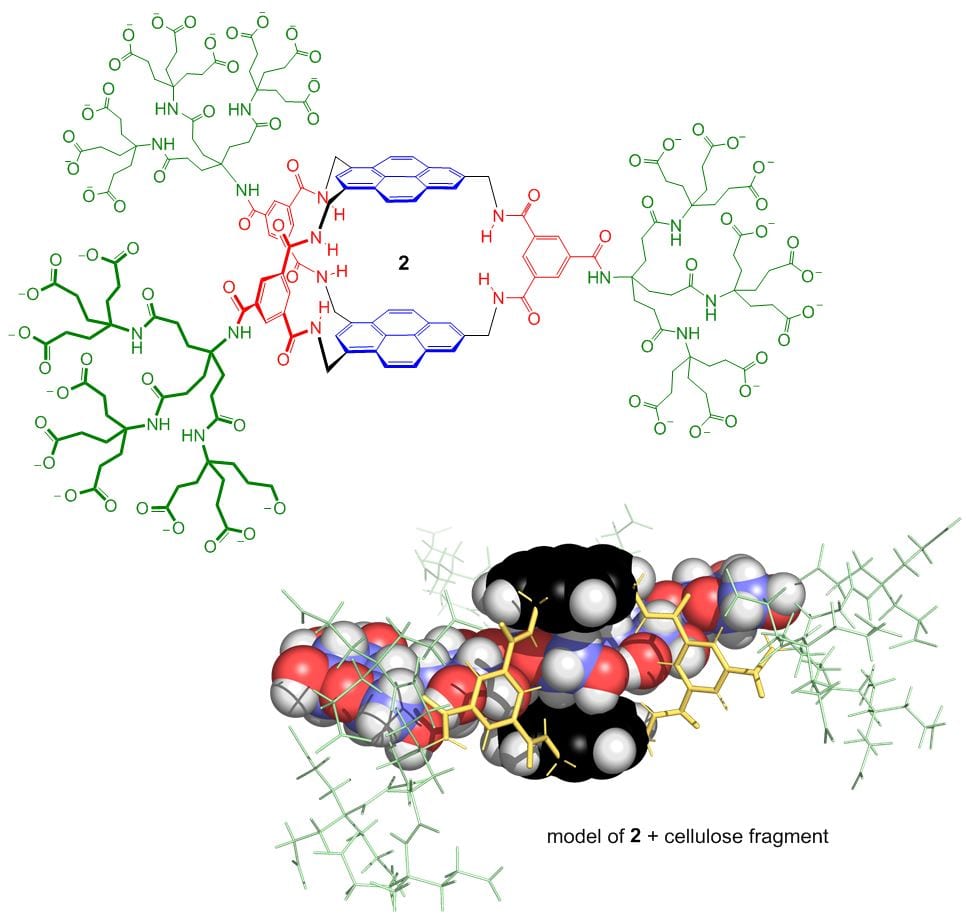
Edvotek Affinity Chromatography of Glucose-Binding Proteins Affinity Chromatography | Fisher Scientific
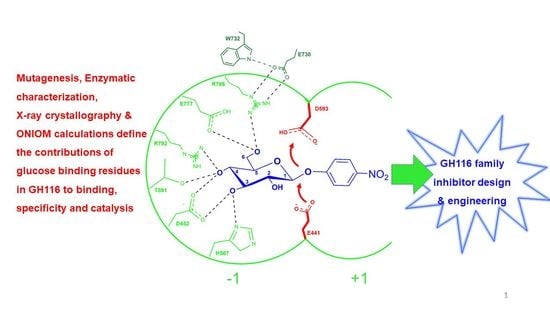
Catalysts | Free Full-Text | Systematic Functional and Computational Analysis of Glucose-Binding Residues in Glycoside Hydrolase Family GH116
Binding interactions of glucose (a) with active site residues of GLUT1... | Download Scientific Diagram
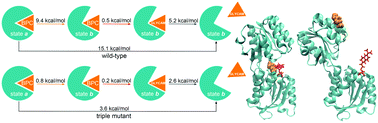
Unravelling the mechanism of glucose binding in a protein-based fluorescence probe: molecular dynamics simulation with a tailor-made charge model - Physical Chemistry Chemical Physics (RSC Publishing)

Sensors | Free Full-Text | D-galactose/D-glucose-binding Protein from Escherichia coli as Probe for a Non-consuming Glucose Implantable Fluorescence Biosensor
Molecular Dynamics Simulation Studies of GLUT4: Substrate-Free and Substrate-Induced Dynamics and ATP-Mediated Glucose Transport Inhibition | PLOS ONE
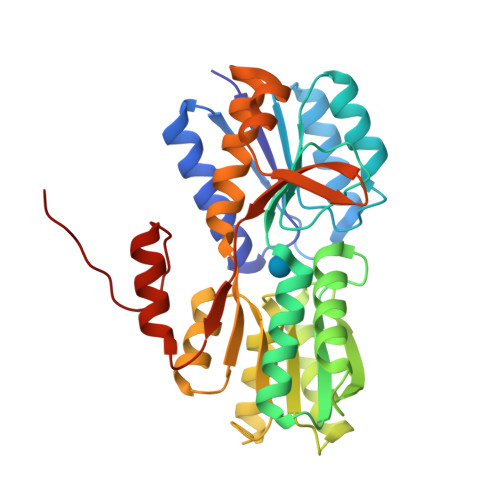
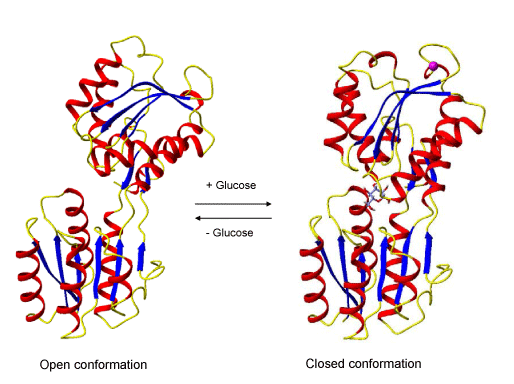
RCSB PDB - 2H3H: Crystal structure of the liganded form of Thermotoga maritima glucose binding protein

Equilibrium binding of glucose to the glucose indicating polymer and... | Download Scientific Diagram

RCSB PDB - 5DVI: High resolution crystal Structure of glucose complexed periplasmic glucose binding protein (ppGBP) from P. putida CSV86